Human microbiota
From Wikipedia, the free encyclopedia
Jump to navigationJump to search
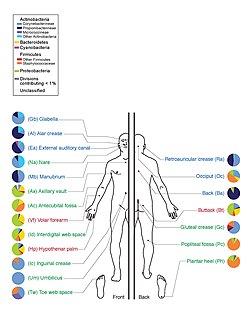
Graphic depicting the human skin microbiota, with relative prevalences of various classes of bacteria
----
The human microbiota is the aggregate of microorganisms that resides on or within any of a number of human tissues and biofluids, including the skin, mammary glands, placenta, seminal fluid, uterus, ovarian follicles, lung, saliva, oral mucosa, conjunctiva, biliary and gastrointestinal tracts. They include bacteria, archaea, fungi, protists and viruses. Though micro-animalscan also live on the human body, they are typically excluded from this definition.
----
The human microbiota is the aggregate of microorganisms that resides on or within any of a number of human tissues and biofluids, including the skin, mammary glands, placenta, seminal fluid, uterus, ovarian follicles, lung, saliva, oral mucosa, conjunctiva, biliary and gastrointestinal tracts. They include bacteria, archaea, fungi, protists and viruses. Though micro-animalscan also live on the human body, they are typically excluded from this definition.
The human microbiome refers specifically to the collective genomes of resident microorganisms.[1]
Humans are colonized by many microorganisms; the traditional estimate is that the average human body is inhabited by ten times as many non-human cells as human cells, but more recent estimates have lowered that ratio to 3:1 or even to approximately the same number.[2][3][4][5] Some microorganisms that colonize humans are commensal, meaning they co-exist without harming humans; others have a mutualistic relationship with their human hosts.[1]:700[6]Conversely, some non-pathogenic microorganisms can harm human hosts via the metabolites they produce, like trimethylamine, which the human body converts to trimethylamine N-oxide via FMO3-mediated oxidation.[7][8] Certain microorganisms perform tasks that are known to be useful to the human host but the role of most of them is not well understood. Those that are expected to be present, and that under normal circumstances do not cause disease, are sometimes deemed normal flora or normal microbiota.[1]
The Human Microbiome Project took on the project of sequencing the genome of the human microbiota, focusing particularly on the microbiota that normally inhabit the skin, mouth, nose, digestive tract, and vagina.[1]
Humans are colonized by many microorganisms; the traditional estimate is that the average human body is inhabited by ten times as many non-human cells as human cells, but more recent estimates have lowered that ratio to 3:1 or even to approximately the same number.[2][3][4][5] Some microorganisms that colonize humans are commensal, meaning they co-exist without harming humans; others have a mutualistic relationship with their human hosts.[1]:700[6]Conversely, some non-pathogenic microorganisms can harm human hosts via the metabolites they produce, like trimethylamine, which the human body converts to trimethylamine N-oxide via FMO3-mediated oxidation.[7][8] Certain microorganisms perform tasks that are known to be useful to the human host but the role of most of them is not well understood. Those that are expected to be present, and that under normal circumstances do not cause disease, are sometimes deemed normal flora or normal microbiota.[1]
The Human Microbiome Project took on the project of sequencing the genome of the human microbiota, focusing particularly on the microbiota that normally inhabit the skin, mouth, nose, digestive tract, and vagina.[1]
It reached a milestone in 2012 when it published its initial results.[9]
Contents
Contents
1Terminology
2Relative numbers
3Study
3.1Shotgun Sequencing
3.1.1Collection of samples and DNA extraction
3.1.2Preparation of the library and sequencing
3.1.3Metagenome assembly
3.1.4Contig binning
3.1.5Analysis after the processing
3.2Marker gene analysis
3.3Phylogenetic Analysis
4Types
4.1Bacteria
4.2Archaea
4.3Fungi
4.4Viruses
5Anatomical areas
5.1Skin
5.2Conjunctiva
5.3Gut
5.4Urethra and bladder
5.5Vagina
5.6Placenta
5.7Uterus
5.8Oral cavity
5.9Lung
5.10Biliary tract
6Disease and death
6.1Cancer
6.2Inflammatory bowel disease
6.3Human immunodeficiency virus
6.4Death
7Environmental health
8Migration
9See also
10Bibliography
11References
12External links
Terminology[edit]
Life timeline
view • discuss • edit
-4500 —
–
-4000 —
–
-3500 —
–
-3000 —
–
-2500 —
–
-2000 —
–
-1500 —
–
-1000 —
–
-500 —
–
0 —
water
Single-celled
life
photosynthesis
Eukaryotes
Multicellular
life
Arthropods Molluscs
Plants
Dinosaurs
Mammals
Flowers
Birds
Primates
←
Earth (−4540)
←
Earliest water
←
Earliest life
←
Earliest oxygen
←
Atmospheric oxygen
←
Oxygen crisis
←
Sexual reproduction
←
Earliest plants
←
Ediacara biota
←
Cambrian explosion
←
Tetrapoda
←
Earliest apes
P
h
a
n
e
r
o
z
o
i
c
P
r
o
t
e
r
o
z
o
i
c
A
r
c
h
e
a
n
H
a
d
e
a
n
Pongola
Huronian
Cryogenian
Andean
Karoo
Quaternary
Ice Ages
(view • discuss)
Axis scale: million years
Also see: Human timeline and Nature timeline
Though widely known as flora ormicroflora, this is a misnomer in technical terms, since the word root flora pertains to plants, and biotarefers to the total collection of organisms in a particular ecosystem. Recently, the more appropriate term microbiota is applied, though its use has not eclipsed the entrenched use and recognition of flora with regard to bacteria and other microorganisms. Both terms are being used in different literature.[6]
Relative numbers[edit]
As of 2014, it was often reported in popular media and in the scientific literature that there are about 10 times as many microbial cells in the human body as there are human cells; this figure was based on estimates that the human microbiome includes around 100 trillion bacterial cells and that an adult human typically has around 10 trillion human cells.[2] In 2014, the American Academy of Microbiologypublished a FAQ that emphasized that the number of microbial cells and the number of human cells are both estimates, and noted that recent research had arrived at a new estimate of the number of human cells – approximately 37.2 trillion, meaning that the ratio of microbial-to-human cells, if the original estimate of 100 trillion bacterial cells is correct, is closer to 3:1.[2][3] In 2016, another group published a new estimate of the ratio being roughly 1:1 (1.3:1, with "an uncertainty of 25% and a variation of 53% over the population of standard 70-kg males").[4][5]
Study[edit]
Main article: Human Microbiome Project

Flowchart illustrating how the human microbiome is studied on the DNA level.
The problem of elucidating the human microbiome is essentially identifying the members of a microbial community which includes bacteria, eukaryotes, and viruses.[10] This is done primarily using DNA-based studies, though RNA, protein and metabolite based studies are also performed.[10][11] DNA-based microbiome studies typically can be categorized as either targeted amplicon studies or more recently shotgunmetagenomic studies. The former focuses on specific known marker genes and is primarily informative taxonomically, while the latter is an entire metagenomic approach which can also be used to study the functional potential of the community.[10] One of the challenges that is present in human microbiome studies, but not in other metagenomic studies is to avoid including the host DNA in the study.[12]
Aside from simply elucidating the composition of the human microbiome, one of the major questions involving the human microbiome is whether there is a "core", that is, whether there is a subset of the community that is shared among most humans.[13][14] If there is a core, then it would be possible to associate certain community compositions with disease states, which is one of the goals of the Human Microbiome Project. It is known that the human microbiome (such as the gut microbiota) is highly variable both within a single subject and among different individuals, a phenomenon which is also observed in mice.[6]
On 13 June 2012, a major milestone of the Human Microbiome Project (HMP) was announced by the NIH director Francis Collins.[9] The announcement was accompanied with a series of coordinated articles published in Nature[15][16] and several journals in the Public Library of Science (PLoS) on the same day. By mapping the normal microbial make-up of healthy humans using genome sequencing techniques, the researchers of the HMP have created a reference database and the boundaries of normal microbial variation in humans. From 242 healthy U.S. volunteers, more than 5,000 samples were collected from tissues from 15 (men) to 18 (women) body sites such as mouth, nose, skin, lower intestine (stool), and vagina. All the DNA, human and microbial, were analyzed with DNA sequencing machines. The microbial genome data were extracted by identifying the bacterial specific ribosomal RNA, 16S rRNA. The researchers calculated that more than 10,000 microbial species occupy the human ecosystem and they have identified 81 – 99% of the genera.
Shotgun Sequencing[edit]
It is frequently difficult to culture in laboratory communities of bacteria, archaea and viruses, therefore sequencing technologies can be exploited in metagenomics, too. Indeed, the complete knowledge of the functions and the characterization of specific microbial strains offer a great potentiality in therapeutic discovery and human health.[17]
Collection of samples and DNA extraction[edit]
The main point is to collect an amount microbial biomass that is sufficient to perform the sequencing and to minimize the sample contamination; for this reason, enrichment techniques can be used. In particular, the DNA extraction method must be good for every bacterial strain, not to have the genomes of the ones that are easy to lyse. Mechanical lysis is usually preferred rather than chemical lysis, and bead beating may result in DNAloss when preparing the library.[17]
Preparation of the library and sequencing[edit]
The most used platforms are Illumina, Ion Torrent, Oxford Nanopore MinION and Pacific Bioscience Sequel, although the Illumina platform is considered the most appealing option due to its wide availability, high output and accuracy. There are no indications regarding the correct amount of sample to use.[17]
Metagenome assembly[edit]
The de novo approach is exploited; however, it presents some difficulties to be overcome. The coverage depends on each genome abundance in its specific community; low-abundance genomes may undergo fragmentation if the sequencing depth is not sufficient enough to avoid the formation of gaps. Luckily, there are metagenome-specific assemblers to help, since, if hundreds of strains are present, the sequencing depth needs to be increased to its maximum.[17]
Contig binning[edit]
Neither from which genome every contig derives, nor the number of genomes present in the sample are known a priori; the aim of this step is to divide the contigs into species. The methods to perform such analysis can be either supervised (database with known sequences) or unsupervised (direct search for contig groups in the collected data). However, both methods require a kind of metric to define a score for the similarity between a specific contig and the group in which it must be put, and algorithms to convert the similarities into allocations in the groups.[17]
Analysis after the processing[edit]
The statistical analysis is essential to validate the obtained results (ANOVA can be used to size the differences between the groups); if it is paired with graphical tools, the outcome is easily visualized and understood.[17]
Once a metagenome is assembled, it is possible to infer the functional potential of the microbiome. The computational challenges for this type of analysis are greater than for single genomes, due the fact that usually metagenomes assemblers have poorer quality, and many recovered genes are non-complete or fragmented. After the gene identification step, the data can be used to carry out a functional annotation by means of multiple alignment of the target genes against orthologs databases.[18]
Marker gene analysis[edit]
It is a technique that exploits primers to target a specific genetic region and enables to determine the microbial phylogenies. The genetic region is characterized by a highly variable region which can confer detailed identification; it is delimited by conserved regions, which function as binding sites for primers used in PCR. The main gene used to characterize bacteria and archaea is 16S rRNA gene, while fungi identification is based on Internal Transcribed Spacer (ITS). The technique is fast and not so expensive and enables to obtain a low-resolution classification of a microbial sample; it is optimal for samples that may be contaminated by host DNA. Primer affinity varies among all DNA sequences, which may result in biases during the amplification reaction; indeed, low-abundance samples are susceptible to overamplification errors, since the other contaminating microorganisms result to be over-represented in case of increasing the PCR cycles. Therefore, the optimization of primer selection can help to decrease such errors, although it requires complete knowledge of the microorganisms present in the sample, and their relative abundances.[19]
Marker gene analysis can be influenced by the primer choice; in this kind of analysis it's desirable to use a well-validated protocol (such as the one used in the Earth Microbiome Project). The first thing to do in a marker gene amplicon analysis is to remove sequencing errors; a lot of sequencing platforms are very reliable, but most of the apparent sequence diversity is still due to errors during the sequencing process. To reduce this phenomenon a first approach is to cluster sequences into Operational taxonomic unit (OTUs): this process consolidates similar sequences (a 97% similarity threshold is usually adopted) into a single feature that can be used in further analysis steps; this method however would discard SNPs because they would get clustered into a single OTU. Another approach is Oligotyping, which includes position-specific information from 16s rRNA sequencing to detect small nucleotide variations and from discriminating between closely related distinct taxa. These methods give as an output a table of DNA sequences and counts of the different sequences per sample rather than OTU.[19]
Another important step in the analysis is to assign a taxonomic name to microbial sequences in the data. This can be done using machine learning approaches that can reach an accuracy at genus-level of about 80%. Other popular analysis packages provide support for taxonomic classification using exact matches to reference databases and should provide greater specificity, but poor sensitivity. Unclassified microorganism should be further checked for organelle sequences.[19]
Phylogenetic Analysis[edit]
Many methods that exploit phylogenetic inference use the 16SRNA gene for Archea and Bacteria and the 18SRNA gene for Eukariotes. Phylogenetic comparative methods (PCS) are based on the comparison of multiple traits among microorganisms; the principle is: the closely they are related, the higher number of traits they share. Usually PCS are coupled with phylogenetic generalized least square (PGLS) or other statistical analysis to get more significant results. Ancestral state reconstruction is used in microbiome studies to impute trait values for taxa whose traits are unknown. This is commonly performed with PICRUSt, which relies on available databases. Phylogenetic variables are chosen by researchers according to the type of study: through the selection of some variables with significant biological informations, it is possible to reduce the dimension of the data to analyse.[20]
Phylogenetic aware distance is usually performed with UniFrac or similar tools, such as Soresen's index or Rao's D, to quantify the differences between the different communities. All this methods are negatively affected by horizontal gene trasmission (HGT), since it can generate errors and lead to the correlation of distant species. There are different ways to reduce the negative impact of HGT: the use of multiple genes or computational tools to assess the probability of putative HGT events.[20]
Types[edit]
Bacteria[edit]
Populations of microbes (such as bacteria and yeasts) inhabit the skin and mucosal surfaces in various parts of the body. Their role forms part of normal, healthy human physiology, however if microbe numbers grow beyond their typical ranges (often due to a compromised immune system) or if microbes populate (such as through poor hygiene or injury) areas of the body normally not colonized or sterile (such as the blood, or the lower respiratory tract, or the abdominal cavity), disease can result (causing, respectively, bacteremia/sepsis, pneumonia, and peritonitis).[medical citation needed]
The Human Microbiome Project found that individuals host thousands of bacterial types, different body sites having their own distinctive communities. Skin and vaginal sites showed smaller diversity than the mouth and gut, these showing the greatest richness. The bacterial makeup for a given site on a body varies from person to person, not only in type, but also in abundance. Bacteria of the same species found throughout the mouth are of multiple subtypes, preferring to inhabit distinctly different locations in the mouth. Even the enterotypes in the human gut, previously thought to be well understood, are from a broad spectrum of communities with blurred taxon boundaries.[21][22]
It is estimated that 500 to 1,000 species of bacteria live in the human gut but belong to just a few phyla: Firmicutes and Bacteroidetes dominate but there are also Proteobacteria, Verrumicrobia, Actinobacteria, Fusobacteria and Cyanobacteria.[23]
A number of types of bacteria, such as Actinomyces viscosus and A. naeslundii, live in the mouth, where they are part of a sticky substance called plaque. If this is not removed by brushing, it hardens into calculus (also called tartar). The same bacteria also secrete acids that dissolve tooth enamel, causing tooth decay.
The vaginal microflora consist mostly of various lactobacillus species. It was long thought that the most common of these species was Lactobacillus acidophilus, but it has later been shown that L. iners is in fact most common, followed by L. crispatus. Other lactobacilli found in the vagina are L. jensenii, L. delbruekii and L. gasseri. Disturbance of the vaginal flora can lead to infections such as bacterial vaginosis or candidiasis ("yeast infection").
Archaea[edit]
Archaea are present in the human gut, but, in contrast to the enormous variety of bacteriain this organ, the numbers of archaeal species are much more limited.[24] The dominant group are the methanogens, particularly Methanobrevibacter smithii and Methanosphaera stadtmanae.[25] However, colonization by methanogens is variable, and only about 50% of humans have easily detectable populations of these organisms.[26]
As of 2007, no clear examples of archaeal pathogens were known,[27][28] although a relationship has been proposed between the presence of some methanogens and human periodontal disease.[29]
Fungi[edit]
See also: Mycobiota (human)
Fungi, in particular yeasts, are present in the human gut.[30][31][32][33] The best-studied of these are Candida species due to their ability to become pathogenic in immunocompromised and even in healthy hosts.[31][32][33] Yeasts are also present on the skin,[30] such as Malassezia species, where they consume oils secreted from the sebaceous glands.[34][35]
Viruses[edit]
See also: Human virome
Viruses, especially bacterial viruses (bacteriophages), colonize various body sites. These colonized sites include the skin,[36] gut,[37] lungs,[38] and oral cavity.[39] Virus communities have been associated with some diseases, and do not simply reflect the bacterial communities.[40][41][42]
Anatomical areas[edit]
Main article: List of human flora
Skin[edit]
Main article: Skin flora
A study of twenty skin sites on each of ten healthy humans found 205 identified genera in nineteen bacterial phyla, with most sequences assigned to four phyla: Actinobacteria(51.8%), Firmicutes (24.4%), Proteobacteria (16.5%), and Bacteroidetes (6.3%).[43] A large number of fungal genera are present on healthy human skin, with some variability by region of the body; however, during pathological conditions, certain genera tend to dominate in the affected region.[30] For example, Malassezia is dominant in atopic dermatitis and Acremonium is dominant on dandruff-afflicted scalps.[30]
The skin acts as a barrier to deter the invasion of pathogenic microbes. The human skin contains microbes that reside either in or on the skin and can be residential or transient. Resident microorganism types vary in relation to skin type on the human body. A majority of microbes reside on superficial cells on the skin or prefer to associate with glands. These glands such as oil or sweat glands provide the microbes with water, amino acids, and fatty acids. In addition, resident bacteria that associated with oil glands are often Gram-positive and can be pathogenic.[1]
Conjunctiva[edit]
A small number of bacteria and fungi are normally present in the conjunctiva.[30][44]Classes of bacteria include Gram-positive cocci (e.g., Staphylococcus and Streptococcus) and Gram-negative rods and cocci (e.g., Haemophilus and Neisseria) are present.[44]Fungal genera include Candida, Aspergillus, and Penicillium.[30] The lachrymal glandscontinuously secrete, keeping the conjunctiva moist, while intermittent blinking lubricates the conjunctiva and washes away foreign material. Tears contain bactericides such as lysozyme, so that microorganisms have difficulty in surviving the lysozyme and settling on the epithelial surfaces.
Gut[edit]
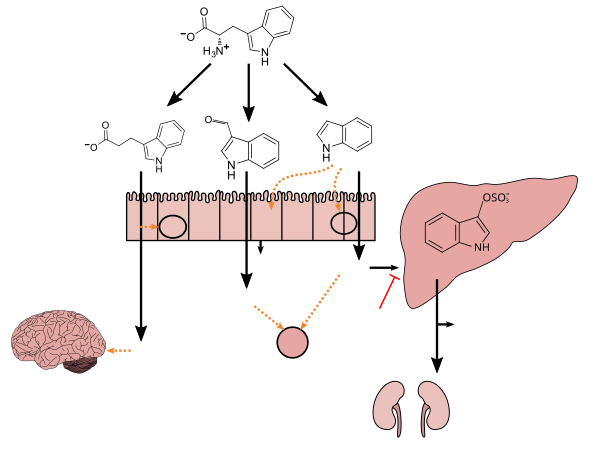
Tryptophan metabolism by human gastrointestinal microbiota (
v
t
e)
Tryptophan
Clostridium
sporogenes
Lacto-
bacilli
Tryptophanase-
expressing
bacteria
IPA
I3A
Indole
Liver
Brain
IPA
I3A
Indole
Indoxyl
sulfate
AST-120
AhR
Intestinal
immune
cells
Intestinal
epithelium
PXR
Mucosal homeostasis:
↓TNF-α
↑Junction protein-
coding mRNAs
L cell
GLP-1
T J
Neuroprotectant:
↓Activation of glial cells and astrocytes
↓4-Hydroxy-2-nonenal levels
↓DNA damage
–Antioxidant
–Inhibits β-amyloid fibril formation
Maintains mucosal reactivity:
↑IL-22 production
Associated with vascular disease:
↑Oxidative stress
↑Smooth muscle cellproliferation
↑Aortic wall thickness and calcification
Associated with chronic kidney disease:
↑Renal dysfunction
–Uremic toxin
Kidneys
Main article: Gut flora
See also: Gut–brain axis
In humans the composition of gut flora is established during birth.[49] Birth by Cesarean section or vaginal delivery also influences the gut's microbial composition. Babies born through the vaginal canal have non-pathogenic, beneficial gut microbiota similar to those found in the mother.[50] However, the gut microbiota of babies delivered by C-section harbors more pathogenic bacteria such as Escherichia coli and Staphylococcus and it takes longer to develop non-pathogenic, beneficial gut microbiota.[51]
The relationship between some gut flora and humans is not merely commensal (a non-harmful coexistence), but rather a mutualistic relationship.[1] Some human gut microorganisms benefit the host by fermentating dietary fiber into short-chain fatty acids(SCFAs), such as acetic acid and butyric acid, which are then absorbed by the host.[6][52]Intestinal bacteria also play a role in synthesizing vitamin B and vitamin K as well as metabolizing bile acids, sterols, and xenobiotics.[1][52] The systemic importance of the SCFAs and other compounds they produce are like hormones and the gut flora itself appears to function like an endocrine organ,[52] and dysregulation of the gut flora has been correlated with a host of inflammatory and autoimmune conditions.[6][53]
The composition of human gut flora changes over time, when the diet changes, and as overall health changes.[6][53] A systematic review of 15 human randomized controlled trialsfrom July 2016 found that certain commercially available strains of probiotic bacteria from the Bifidobacterium and Lactobacillus genera (B. longum, B. breve, B. infantis, L. helveticus, L. rhamnosus, L. plantarum, and L. casei), when taken by mouth in daily doses of 109–1010 colony forming units (CFU) for 1–2 months, possess treatment efficacy (i.e., improves behavioral outcomes) in certain central nervous system disorders – including anxiety, depression, autism spectrum disorder, and obsessive–compulsive disorder – and improves certain aspects of memory.[54] However, changes in the composition of gut microbiota has also been found to be correlated with harmful effects on health. In an article published by Musso et al., it was found that the gut microbiota of obese invidividuals had more Firmicutes and less Bacteroidetes than healthy individuals.[55] Furthermore, a study done by Gordon et al., confirmed that it was the composition of the microbiota that causes obesity rather than the other way around. This was done by transplanting the gut microbiota from diet-induced obese(DIO) mice or lean control mice into lean germ-free mice that do not have a microbiome. They found that the mice transplanted with DIO mouse gut microbiota had significantly higher total body fat than the mice transplanted with lean mouse microbiota when fed with the same diet.[56]
Urethra and bladder[edit]
The genitourinary system appears to have a microbiota,[57][58] which is an unexpected finding in light of the long-standing use of standard clinical microbiological culture methods to detect bacteria in urine when people show signs of a urinary tract infection; it is common for these tests to show no bacteria present.[59] It appears that common culture methods do not detect many kinds of bacteria and other microorganisms that are normally present.[59]As of 2017, sequencing methods were used to identify these microorganisms to determine if there are differences in microbiota between people with urinary tract problems and those who are healthy.[57][58]
Vagina[edit]
Main article: Vaginal flora
See also: List of microbiota species of the lower reproductive tract of women, List of bacterial vaginosis microbiota, and Vaginal microbiota in pregnancy
Vaginal microbiota refers to those species and genera that colonize the vagina. These organisms play an important role in protecting against infections and maintaining vaginal health.[60] The most abundant vaginal microorganisms found in premenopausal women are from the genus Lactobacillus, which suppress pathogens by producing hydrogen peroxide and lactic acid.[32][60][61] Bacterial species composition and ratios vary depending on the stage of the menstrual cycle.[62][63][needs update] Ethnicity also influences vaginal flora. The occurrence of hydrogen peroxide-producing lactobacilli is lower in African American women and vaginal pH is higher.[64] Other influential factors such as sexual intercourse and antibiotics have been linked to the loss of lactobacilli.[61] Moreover, studies have found that sexual intercourse with a condom does appear to change lactobacilli levels, and does increase the level of Escherichia coli within the vaginal flora.[61] Changes in the normal, healthy vaginal microbiota is an indication of infections, such as candidiasisor bacterial vaginosis.[61] Candida albicans inhibits the growth of Lactobacillus species, while Lactobacillus species which produce hydrogen peroxide inhibit the growth and virulence of Candida albicans in both the vagina and the gut.[30][32][33]
Fungal genera that have been detected in the vagina include Candida, Pichia, Eurotium, Alternaria, Rhodotorula, and Cladosporium, among others.[30]
Placenta[edit]
Main article: Placental microbiome
Until recently the placenta was considered to be a sterile organ but commensal, nonpathogenic bacterial species and genera have been identified that reside in the placental tissue.[65][66][67]
Uterus[edit]
Main article: Uterine microbiome
Until recently, the upper reproductive tract of women was considered to be a sterile environment. A variety of microorganisms inhabit the uterus of healthy, asymptomatic women of reproductive age. The microbiome of the uterus differs significantly from that of the vagina and gastrointestinal tract.[68]
Oral cavity[edit]
Main article: Oral microbiology
The environment present in the human mouth allows the growth of characteristic microorganisms found there. It provides a source of water and nutrients, as well as a moderate temperature.[1] Resident microbes of the mouth adhere to the teeth and gums to resist mechanical flushing from the mouth to stomach where acid-sensitive microbes are destroyed by hydrochloric acid.[1][32]
Anaerobic bacteria in the oral cavity include: Actinomyces, Arachnia, Bacteroides, Bifidobacterium, Eubacterium, Fusobacterium, Lactobacillus, Leptotrichia, Peptococcus, Peptostreptococcus, Propionibacterium, Selenomonas, Treponema, and Veillonella.[69][needs update] Genera of fungi that are frequently found in the mouth include Candida, Cladosporium, Aspergillus, Fusarium, Glomus, Alternaria, Penicillium, and Cryptococcus, among others.[30]
Bacteria accumulate on both the hard and soft oral tissues in biofilm allowing them to adhere and strive in the oral environment while protected from the environmental factors and antimicrobial agents.[70] Saliva plays a key biofilm homeostatic role allowing recolonization of bacteria for formation and controlling growth by detaching biofilm buildup.[71] It also provides a means of nutrients and temperature regulation. The location of the biofilm determines the type of exposed nutrients it receives.[72]
Oral bacteria have evolved mechanisms to sense their environment and evade or modify the host. However, a highly efficient innate host defense system constantly monitors the bacterial colonization and prevents bacterial invasion of local tissues. A dynamic equilibrium exists between dental plaque bacteria and the innate host defense system.[73]
This dynamic between host oral cavity and oral microbes plays a key role in health and disease as it provides entry into the body.[74] A healthy equilibrium presents a symbiotic relationship where oral microbes limit growth and adherence of pathogens while the host provides an environment for them to flourish.[74][70] Ecological changes such as change of immune status, shift of resident microbes and nutrient availability shift from a mutual to parasitic relationship resulting in the host being prone to oral and systemic disease.[70]Systemic diseases such as diabetes and cardiovascular diseases has been correlated to poor oral health.[74] Of particular interest is the role of oral microorganisms in the two major dental diseases: dental caries and periodontal disease.[73] Pathogen colonization at the periodontium cause an excessive immune response resulting in a periodontal pocket- a deepened space between the tooth and gingiva.[70] This acts as a protected blood-rich reservoir with nutrients for anaerobic pathogens.[70] Systemic disease at various sites of the body can result from oral microbes entering the blood bypassing periodontal pockets and oral membranes.[74]
Persistent proper oral hygiene is the primary method for preventing oral and systemic disease.[74] It reduces the density of biofilm and overgrowth of potential pathogenic bacteria resulting in disease.[72] However, proper oral hygiene may not be enough as the oral microbiome, genetics, and changes to immune response play a factor in developing chronic infections.[72] Use of antibiotics could treat already spreading infection but ineffective against bacteria within biofilms.[72]
Lung[edit]
Main article: Lung microbiome
Much like the oral cavity, the upper and lower respiratory system possess mechanical deterrents to remove microbes. Goblet cells produce mucous which traps microbes and moves them out of the respiratory system via continuously moving ciliated epithelial cells.[1] In addition, a bactericidal effect is generated by nasal mucus which contains the enzyme lysozyme.[1] The upper and lower respiratory tract appears to have its own set of microbiota.[75] Pulmonary bacterial microbiota belong to 9 major bacterial genera: Prevotella, Sphingomonas, Pseudomonas, Acinetobacter, Fusobacterium, Megasphaera, Veillonella, Staphylococcus, and Streptococcus. Some of the bacteria considered "normal biota" in the respiratory tract can cause serious disease especially in immunocompromised individuals; these include Streptococcus pyogenes, Haemophilus influenzae, Streptococcus pneumoniae, Neisseria meningitidis, and Staphylococcus aureus.[citation needed] Fungal genera that compose the pulmonary mycobiome include Candida, Malassezia, Neosartorya, Saccharomyces, and Aspergillus, among others.[30]
Unusual distributions of bacterial and fungal genera in the respiratory tract is observed in people with cystic fibrosis.[30][76] Their bacterial flora often contains antibiotic-resistant and slow-growing bacteria, and the frequency of these pathogens changes in relation to age.[76]
Biliary tract[edit]
Traditionally the biliary tract has been considered to be normally sterile, and the presence of microorganisms in bile is a marker of pathological process. This assumption was confirmed by failure in allocation of bacterial strains from the normal bile duct. Papers began emerging in 2013 showing that the normal biliary microbiota is a separate functional layer which protects a biliary tract from colonization by exogenous microorganisms.[77]
Disease and death[edit]
A symbiotic relationship between the gut microbiota and different bacteria may influence an individual's immune response.[78]
Cancer[edit]
Although cancer is generally a disease of host genetics and environmental factors, microorganisms are implicated in some 20% of human cancers.[79] Particularly for potential factors in colon cancer, bacterial density is one million times higher than in the small intestine, and approximately 12-fold more cancers occur in the colon compared to the small intestine, possibly establishing a pathogenic role for microbiota in colon and rectal cancers.[80] Microbial density may be used as a prognostic tool in assessment of colorectal cancers.[80]
The microbiota may affect carcinogenesis in three broad ways: (i) altering the balance of tumor cell proliferation and death, (ii) regulating immune system function, and (iii) influencing metabolism of host-produced factors, foods and pharmaceuticals.[79] Tumors arising at boundary surfaces, such as the skin, oropharynx and respiratory, digestive and urogenital tracts, harbor a microbiota. Substantial microbe presence at a tumor site does not establish association or causal links. Instead, microbes may find tumor oxygen tensionor nutrient profile supportive. Decreased populations of specific microbes or induced oxidative stress may also increase risks.[79][80] Of the around 1030 microbes on earth, ten are designated by the International Agency for Research on Cancer as human carcinogens.[79] Microbes may secrete proteins or other factors directly drive cell proliferation in the host, or may up- or down-regulate the host immune system including driving acute or chronic inflammation in ways that contribute to carcinogenesis.[79]
Concerning the relationship of immune function and development of inflammation, mucosal surface barriers are subject to environmental risks and must rapidly repair to maintain homeostasis. Compromised host or microbiota resiliency also reduce resistance to malignancy, possibly inducing inflammation and cancer. Once barriers are breached, microbes can elicit proinflammatory or immunosuppressive programs through various pathways.[79] For example, cancer-associated microbes appear to activate NF-κΒ signaling within the tumor microenviroment. Other pattern recognition receptors, such as nucleotide-binding oligomerization domain–like receptor (NLR) family members NOD-2, NLRP3, NLRP6 and NLRP12, may play a role in mediating colorectal cancer.[79] Likewise Helicobacter pylori appears to increase the risk of gastric cancer, due to its driving a chronic inflammatory response in the stomach.[79][80]
Inflammatory bowel disease[edit]
Inflammatory bowel disease consists of two different diseases: ulcerative colitis and Crohn's disease and both of these diseases present with disruptions in the gut microbiota (also known as dysbiosis). This dysbiosis presents itself in the form of decreased microbial diversity in the gut,[81][82] and is correlated to defects in host genes that changes the innate immune response in individuals.[81]
Human immunodeficiency virus[edit]
The HIV disease progression influences the composition and function of the gut microbiota, with notable differences between HIV-negative, HIV-positive, and post-ARTHIV-positive populations.[citation needed] HIV decreases the integrity of the gut epithelial barrier function by affecting tight junctions. This breakdown allows for translocation across the gut epithelium, which is thought to contribute to increases in inflammation seen in people with HIV.[83]
Vaginal microbiota plays a role in the infectivity of HIV, with an increased risk of infection and transmission when the woman has bacterial vaginosis, a condition characterized by an abnormal balance of vaginal bacteria.[84] The enhanced infectivity is seen with the increase in pro-inflammatory cytokines and CCR5 + CD4+ cells in the vagina. However, a decrease in infectivity is seen with increased levels of vaginal Lactobacillus, which promotes an anti-inflammatory condition.[83]
Death[edit]
Main article: necrobiome
With death the microbiome of the living body collapses and a different microbiome named necrobiome establishes itself. Its predictable changes over time are thought to be useful to help determine the time of death.[85][86]
Environmental health[edit]
Studies in 2009 questioned whether the decline in biota (including microfauna) as a result of human intervention might impede human health, hospital safety procedures, food product design, and treatments of disease.[87]
Migration[edit]
Preliminary research indicates that immediate changes in the microbiota may occur when a person migrates from one country to another, such as when Thai immigrants settled in the United States.[88] Losses of microbiota diversity were greater in obese individuals and children of immigrants.[88]
See also[edit]
Drug resistance
Human Microbiome Project
Human milk microbiome
Human virome
Initial acquisition of microbiota
List of bacterial vaginosis microbiota
Microbiome
Microorganism
uBiome
Microbiome Immunity Project
Carbon monoxide-releasing molecules
---------
Bibliography[edit]
Ed Yong. I Contain Multitudes: The Microbes Within Us and a Grander View of Life.368 pages, Published 9 August 2016 by Ecco, ISBN 0062368591.
References[edit]
- ^ Jump up to:a b c d e f g h i j k Sherwood, Linda; Willey, Joanne; Woolverton, Christopher (2013). Prescott's Microbiology (9th ed.). New York: McGraw Hill. pp. 713–721. ISBN 9780073402406. OCLC 886600661.
- ^ Jump up to:a b c American Academy of Microbiology FAQ: Human Microbiome January 2014
- ^ Jump up to:a b Judah L. Rosner for Microbe Magazine, February 2014. Ten Times More Microbial Cells than Body Cells in Humans?
- ^ Jump up to:a b Alison Abbott for Nature News. 8 January 2016 Scientists bust myth that our bodies have more bacteria than human cells
- ^ Jump up to:a b Sender, R; Fuchs, S; Milo, R (January 2016). "Are We Really Vastly Outnumbered? Revisiting the Ratio of Bacterial to Host Cells in Humans". Cell. 164 (3): 337–40. doi:10.1016/j.cell.2016.01.013. PMID 26824647.
- ^ Jump up to:a b c d e f Quigley, EM (2013). "Gut bacteria in health and disease". Gastroenterol Hepatol (N Y). 9 (9): 560–9. PMC 3983973. PMID 24729765.
- ^ Falony G, Vieira-Silva S, Raes J (2015). "Microbiology Meets Big Data: The Case of Gut Microbiota-Derived Trimethylamine". Annu. Rev. Microbiol. 69: 305–321. doi:10.1146/annurev-micro-091014-104422. PMID 26274026. we review literature on trimethylamine (TMA), a microbiota-generated metabolite linked to atherosclerosis development.
- ^ Gaci N, Borrel G, Tottey W, O'Toole PW, Brugère JF (November 2014). "Archaea and the human gut: new beginning of an old story". World J. Gastroenterol. 20 (43): 16062–16078. doi:10.3748/wjg.v20.i43.16062. PMC 4239492. PMID 25473158. Trimethylamine is exclusively a microbiota-derived product of nutrients (lecithin, choline, TMAO, L-carnitine) from normal diet, from which seems originate two diseases, trimethylaminuria (or Fish-Odor Syndrome) and cardiovascular disease through the proatherogenic property of its oxidized liver-derived form.
- ^ Jump up to:a b "NIH Human Microbiome Project defines normal bacterial makeup of the body". NIH News. 13 June 2012.
- ^ Jump up to:a b c NIH Human Microbiome Working Group (2009). "The NIH Human Microbiome Project". Genome Res. 19 (12): 2317–2323. doi:10.1101/gr.096651.109. PMC 2792171. PMID 19819907.
- ^ Kuczynski, J.; et al. (2011). "Experimental and analytical tools for studying the human microbiome". Nature Reviews Genetics. 13 (1): 47–58. doi:10.1038/nrg3129. PMC 5119550. PMID 22179717.
- ^ Vestheim, H.; Jarman, S. N. (2008). "Blocking primers to enhance PCR amplification of rare sequences in mixed samples – a case study on prey DNA in Antarctic krill stomachs". Frontiers in Zoology. 5: 12. doi:10.1186/1742-9994-5-12. PMC 2517594. PMID 18638418.
- ^ Tap, Julien; Mondot; et al. (2009). "Towards the human intestinal microbiota phylogenetic core". Environmental Microbiology. 11 (102): 2574–2584. doi:10.1111/j.1462-2920.2009.01982.x. PMID 19601958.
- ^ Hamady, M.; Knight, R. (2009). "Microbial community profiling for human microbiome projects: Tools, techniques, and challenges". Genome Research. 19 (7): 1141–1152. doi:10.1101/gr.085464.108. PMID 19383763.
- ^ Human Microbiome Project Consortium (2012). "A framework for human microbiome research". Nature. 486 (7402): 215–221. Bibcode:2012Natur.486..215T. doi:10.1038/nature11209. PMC 3377744. PMID 22699610.
- ^ The Human Microbiome Project Consortium (2012). "Structure, function and diversity of the healthy human microbiome". Nature. 486 (7402): 207–214. Bibcode:2012Natur.486..207T. doi:10.1038/nature11234. PMC 3564958. PMID 22699609.
- ^ Jump up to:a b c d e f Quince, Christopher; Walker, Alan W; Simpson, Jared T; Loman, Nicholas J; Segata, Nicola (2017-09-12). "Shotgun metagenomics, from sampling to analysis". Nature Biotechnology. 35 (9): 833–844. doi:10.1038/nbt.3935. ISSN 1087-0156.
- ^ Claesson, Marcus J.; Clooney, Adam G.; O'Toole, Paul W. (2017-08-09). "A clinician's guide to microbiome analysis". Nature Reviews Gastroenterology & Hepatology. 14 (10). doi:10.1038/nrgastro.2017.97. ISSN 1759-5045.
- ^ Jump up to:a b c Knight, Rob; Vrbanac, Alison; Taylor, Bryn C.; Aksenov, Alexander; Callewaert, Chris; Debelius, Justine; Gonzalez, Antonio; Kosciolek, Tomasz; McCall, Laura-Isobel (2018-05-23). "Best practices for analysing microbiomes". Nature Reviews Microbiology. 16 (7): 410–422. doi:10.1038/s41579-018-0029-9. ISSN 1740-1526.
- ^ Jump up to:a b Washburne, Alex D.; Morton, James T.; Sanders, Jon; McDonald, Daniel; Zhu, Qiyun; Oliverio, Angela M.; Knight, Rob (2018-05-24). "Methods for phylogenetic analysis of microbiome data". Nature Microbiology. 3 (6): 652–661. doi:10.1038/s41564-018-0156-0. ISSN 2058-5276.
- ^ PLoS Human Microbiome Project Collection Manuscript Summaries 13 June 2012
- ^ "Consortium of Scientists Map the Human Body's Bacterial Ecosystem". ucsf.edu.
- ^ Sommer, F; Bäckhed, F (April 2013). "The gut microbiota—masters of host development and physiology". Nat Rev Microbiol. 11 (4): 227–38. doi:10.1038/nrmicro2974. PMID 23435359.
- ^ Eckburg PB, Bik EM, Bernstein CN, et al. (2005). "Diversity of the human intestinal microbial flora". Science. 308 (5728): 1635–8. Bibcode:2005Sci...308.1635E. doi:10.1126/science.1110591. PMC 1395357. PMID 15831718.
- ^ Duncan SH, Louis P, Flint HJ (2007). "Cultivable bacterial diversity from the human colon". Lett. Appl. Microbiol. 44 (4): 343–50. doi:10.1111/j.1472-765X.2007.02129.x. PMID 17397470.
- ^ Florin TH, Zhu G, Kirk KM, Martin NG (2000). "Shared and unique environmental factors determine the ecology of methanogens in humans and rats". Am. J. Gastroenterol. 95 (10): 2872–9. doi:10.1111/j.1572-0241.2000.02319.x. PMID 11051362.
- ^ Eckburg P, Lepp P, Relman D (2003). "Archaea and their potential role in human disease". Infection and Immunity. 71 (2): 591–6. doi:10.1128/IAI.71.2.591-596.2003. PMC 145348. PMID 12540534.
- ^ Cavicchioli R, Curmi P, Saunders N, Thomas T (2003). "Pathogenic archaea: do they exist?". BioEssays. 25 (11): 1119–28. doi:10.1002/bies.10354. PMID 14579252.
- ^ Lepp P, Brinig M, Ouverney C, Palm K, Armitage G, Relman D (2004). "Methanogenic Archaea and human periodontal disease". Proc. Natl. Acad. Sci. U.S.A. 101 (16): 6176–81. Bibcode:2004PNAS..101.6176L. doi:10.1073/pnas.0308766101. PMC 395942. PMID 15067114.
- ^ Jump up to:a b c d e f g h i j k Cui L, Morris A, Ghedin E (July 2013). "The human mycobiome in health and disease". Genome Med. 5 (7): 63. doi:10.1186/gm467. PMC 3978422. PMID 23899327. Figure 2: Distribution of fungal genera in different body sites
- ^ Jump up to:a b Martins N, Ferreira IC, Barros L, Silva S, Henriques M (June 2014). "Candidiasis: predisposing factors, prevention, diagnosis and alternative treatment". Mycopathologia. 177(5–6): 223–240. doi:10.1007/s11046-014-9749-1. PMID 24789109. Candida species and other microorganisms are involved in this complicated fungal infection, but Candida albicans continues to be the most prevalent. In the past two decades, it has been observed an abnormal overgrowth in the gastrointestinal, urinary and respiratory tracts, not only in immunocompromised patients, but also related to nosocomial infections and even in healthy individuals. There is a widely variety of causal factors that contribute to yeast infection which means that candidiasis is a good example of a multifactorial syndrome.
- ^ Jump up to:a b c d e Wang ZK, Yang YS, Stefka AT, Sun G, Peng LH (April 2014). "Review article: fungal microbiota and digestive diseases". Aliment. Pharmacol. Ther. 39 (8): 751–766. doi:10.1111/apt.12665. PMID 24612332. In addition, GI fungal infection is reported even among those patients with normal immune status. Digestive system-related fungal infections may be induced by both commensal opportunistic fungi and exogenous pathogenic fungi. ... Candida sp. is also the most frequently identified species among patients with gastric IFI. ... It was once believed that gastric acid could kill microbes entering the stomach and that the unique ecological environment of the stomach was not suitable for microbial colonisation or infection. However, several studies using culture-independent methods confirmed that large numbers of acid-resistant bacteria belonging to eight phyla and up to 120 species exist in the stomach, such as Streptococcus sp., Neisseria sp. and Lactobacillus sp. etc.26, 27Furthermore, Candida albicans can grow well in highly acidic environments,28 and some genotypes may increase the severity of gastric mucosal lesions.29
- ^ Jump up to:a b c Erdogan A, Rao SS (April 2015). "Small intestinal fungal overgrowth". Curr Gastroenterol Rep. 17 (4): 16. doi:10.1007/s11894-015-0436-2. PMID 25786900. Small intestinal fungal overgrowth (SIFO) is characterized by the presence of excessive number of fungal organisms in the small intestine associated with gastrointestinal (GI) symptoms. Candidiasis is known to cause GI symptoms particularly in immunocompromised patients or those receiving steroids or antibiotics. However, only recently, there is emerging literature that an overgrowth of fungus in the small intestine of non-immunocompromised subjects may cause unexplained GI symptoms. Two recent studies showed that 26 % (24/94) and 25.3 % (38/150) of a series of patients with unexplained GI symptoms had SIFO. The most common symptoms observed in these patients were belching, bloating, indigestion, nausea, diarrhea, and gas. ... Fungal-bacterial interaction may act in different ways and may either be synergistic or antagonistic or symbiotic [29]. Some bacteria such as Lactobacillus species can interact and inhibit both the virulence and growth of Candida species in the gut by producing hydrogen peroxide [30]. Any damage to the mucosal barrier or disruption of GI microbiota with chemotherapy or antibiotic use, inflammatory processes, activation of immune molecules and disruption of epithelial repair may all cause fungal overgrowth [27].
- ^ Marcon MJ, Powell DA (1 April 1992). "Human infections due to Malassezia spp". Clin. Microbiol. Rev. 5 (2): 101–19. doi:10.1128/CMR.5.2.101. PMC 358230. PMID 1576583.
- ^ Roth RR, James WD (1988). "Microbial ecology of the skin". Annu. Rev. Microbiol. 42 (1): 441–64. doi:10.1146/annurev.mi.42.100188.002301. PMID 3144238.
- ^ Hannigan, Geoffrey D.; Meisel, Jacquelyn S.; Tyldsley, Amanda S.; Zheng, Qi; Hodkinson, Brendan P.; SanMiguel, Adam J.; Minot, Samuel; Bushman, Frederic D.; Grice, Elizabeth A. (30 October 2015). "The Human Skin Double-Stranded DNA Virome: Topographical and Temporal Diversity, Genetic Enrichment, and Dynamic Associations with the Host Microbiome". MBio. 6 (5): e01578–15. doi:10.1128/mBio.01578-15. PMC 4620475. PMID 26489866.
- ^ Minot, Samuel; Sinha, Rohini; Chen, Jun; Li, Hongzhe; Keilbaugh, Sue A.; Wu, Gary D.; Lewis, James D.; Bushman, Frederic D. (1 October 2011). "The human gut virome: inter-individual variation and dynamic response to diet". Genome Research. 21 (10): 1616–1625. doi:10.1101/gr.122705.111. PMC 3202279. PMID 21880779.
- ^ Young, J. C.; Chehoud, C.; Bittinger, K.; Bailey, A.; Diamond, J. M.; Cantu, E.; Haas, A. R.; Abbas, A.; Frye, L. (1 January 2015). "Viral metagenomics reveal blooms of anelloviruses in the respiratory tract of lung transplant recipients". American Journal of Transplantation. 15(1): 200–209. doi:10.1111/ajt.13031. PMC 4276431. PMID 25403800.
- ^ Abeles, Shira R.; Robles-Sikisaka, Refugio; Ly, Melissa; Lum, Andrew G.; Salzman, Julia; Boehm, Tobias K.; Pride, David T. (1 September 2014). "Human oral viruses are personal, persistent and gender-consistent". The ISME Journal. 8 (9): 1753–1767. doi:10.1038/ismej.2014.31. PMC 4139723. PMID 24646696.
- ^ Ly, Melissa; Abeles, Shira R.; Boehm, Tobias K.; Robles-Sikisaka, Refugio; Naidu, Mayuri; Santiago-Rodriguez, Tasha; Pride, David T. (1 July 2014). "Altered Oral Viral Ecology in Association with Periodontal Disease". MBio. 5 (3): e01133–14. doi:10.1128/mBio.01133-14. PMC 4030452. PMID 24846382.
- ^ Monaco, Cynthia L.; Gootenberg, David B.; Zhao, Guoyan; Handley, Scott A.; Ghebremichael, Musie S.; Lim, Efrem S.; Lankowski, Alex; Baldridge, Megan T.; Wilen, Craig B. (9 March 2016). "Altered Virome and Bacterial Microbiome in Human Immunodeficiency Virus-Associated Acquired Immunodeficiency Syndrome". Cell Host & Microbe. 19 (3): 311–322. doi:10.1016/j.chom.2016.02.011. PMC 4821831. PMID 26962942.
- ^ Norman, Jason M.; Handley, Scott A.; Baldridge, Megan T.; Droit, Lindsay; Liu, Catherine Y.; Keller, Brian C.; Kambal, Amal; Monaco, Cynthia L.; Zhao, Guoyan (29 January 2015). "Disease-specific alterations in the enteric virome in inflammatory bowel disease". Cell. 160(3): 447–460. doi:10.1016/j.cell.2015.01.002. PMC 4312520. PMID 25619688.
- ^ Grice, Elizabeth A.; Kong, HH; Conlan, S; Deming, CB; Davis, J; Young, AC; NISC Comparative Sequencing Program; Bouffard, GG; et al. (2006). "Topographical and Temporal Diversity of the Human Skin Microbiome". Science. 324 (5931): 1190–2. Bibcode:2009Sci...324.1190G. doi:10.1126/science.1171700. PMC 2805064. PMID 19478181.
- ^ Jump up to:a b "The Normal Bacterial Flora of Humans". textbookofbacteriology.net.
- ^ Jump up to:a b c d e f g h i Zhang LS, Davies SS (April 2016). "Microbial metabolism of dietary components to bioactive metabolites: opportunities for new therapeutic interventions". Genome Med. 8 (1): 46. doi:10.1186/s13073-016-0296-x. PMC 4840492. PMID 27102537. Lactobacillus spp. convert tryptophan to indole-3-aldehyde (I3A) through unidentified enzymes [125]. Clostridium sporogenes convert tryptophan to IPA [6], likely via a tryptophan deaminase. ... IPA also potently scavenges hydroxyl radicals
- Table 2: Microbial metabolites: their synthesis, mechanisms of action, and effects on health and disease
- Figure 1: Molecular mechanisms of action of indole and its metabolites on host physiology and disease
- ^ Wikoff WR, Anfora AT, Liu J, Schultz PG, Lesley SA, Peters EC, Siuzdak G (March 2009). "Metabolomics analysis reveals large effects of gut microflora on mammalian blood metabolites". Proc. Natl. Acad. Sci. U.S.A. 106 (10): 3698–3703. doi:10.1073/pnas.0812874106. PMC 2656143. PMID 19234110. Production of IPA was shown to be completely dependent on the presence of gut microflora and could be established by colonization with the bacterium Clostridium sporogenes.
- IPA metabolism diagram
- ^ "3-Indolepropionic acid". Human Metabolome Database. University of Alberta. Retrieved 12 June 2018. Indole-3-propionate (IPA), a deamination product of tryptophan formed by symbiotic bacteria in the gastrointestinal tract of mammals and birds. 3-Indolepropionic acid has been shown to prevent oxidative stress and death of primary neurons and neuroblastoma cells exposed to the amyloid beta-protein in the form of amyloid fibrils, one of the most prominent neuropathologic features of Alzheimer's disease. 3-Indolepropionic acid also shows a strong level of neuroprotection in two other paradigms of oxidative stress. (PMID 10419516) ... More recently it has been found that higher indole-3-propionic acid levels in serum/plasma are associated with reduced likelihood of type 2 diabetes and with higher levels of consumption of fiber-rich foods (PMID 28397877)
- Origin: • Endogenous • Microbial
- ^ Chyan YJ, Poeggeler B, Omar RA, Chain DG, Frangione B, Ghiso J, Pappolla MA (July 1999). "Potent neuroprotective properties against the Alzheimer beta-amyloid by an endogenous melatonin-related indole structure, indole-3-propionic acid". J. Biol. Chem. 274(31): 21937–21942. doi:10.1074/jbc.274.31.21937. PMID 10419516. [Indole-3-propionic acid (IPA)] has previously been identified in the plasma and cerebrospinal fluid of humans, but its functions are not known. ... In kinetic competition experiments using free radical-trapping agents, the capacity of IPA to scavenge hydroxyl radicals exceeded that of melatonin, an indoleamine considered to be the most potent naturally occurring scavenger of free radicals. In contrast with other antioxidants, IPA was not converted to reactive intermediates with pro-oxidant activity.
- ^ Yang, Irene; Corwin, Elizabeth J.; Brennan, Patricia A.; Jordan, Sheila; Murphy, Jordan R.; Dunlop, Anne (2016). "The Infant Microbiome". Nursing Research. 65 (1): 76–88. doi:10.1097/NNR.0000000000000133. PMC 4681407. PMID 26657483.
- ^ Mueller, Noel T.; Bakacs, Elizabeth; Combellick, Joan; Grigoryan, Zoya; Dominguez-Bello, Maria G. (February 2015). "The infant microbiome development: mom matters". Trends in Molecular Medicine. 21 (2): 109–117. doi:10.1016/j.molmed.2014.12.002. PMC 4464665. PMID 25578246.
- ^ Wall, R; Ross, R.P; Ryan, C.A; Hussey, S; Murphy, B; Fitzgerald, G.F; Stanton, C (4 March 2009). "Role of Gut Microbiota in Early Infant Development". Clinical Medicine. Pediatrics. 3: 45–54. ISSN 1178-220X. PMC 3676293. PMID 23818794.
- ^ Jump up to:a b c Clarke, G; et al. (August 2014). "Minireview: Gut microbiota: the neglected endocrine organ". Mol Endocrinol. 28 (8): 1221–38. doi:10.1210/me.2014-1108. PMID 24892638.
- ^ Jump up to:a b Shen, S; Wong, CH (2016). "Bugging inflammation: role of the gut microbiota". Clin Transl Immunology. 5 (4): e72. doi:10.1038/cti.2016.12. PMC 4855262. PMID 27195115.
- ^ Wang H, Lee IS, Braun C, Enck P (July 2016). "Effect of probiotics on central nervous system functions in animals and humans – a systematic review". J. Neurogastroenterol Motil. 22 (4): 589–605. doi:10.5056/jnm16018. PMC 5056568. PMID 27413138. These probiotics showed efficacy in improving psychiatric disorder-related behaviors including anxiety, depression, autism spectrum disorder (ASD), obsessive-compulsive disorder, and memory abilities, including spatial and non-spatial memory. Because many of the basic science studies showed some efficacy of probiotics on central nervous system function, this background may guide and promote further preclinical and clinical studies. ... According to the qualitative analyses of current studies, we can provisionally draw the conclusion that B. longum, B. breve, B. infantis, L. helveticus, L. rhamnosus, L. plantarum, and L. casei were most effective in improving CNS function, including psychiatric disease-associated functions (anxiety, depression, mood, stress response) and memory abilities
- ^ Musso, G.; Gambino, R.; Cassader, M. (28 September 2010). "Obesity, Diabetes, and Gut Microbiota: The hygiene hypothesis expanded?". Diabetes Care. 33 (10): 2277–2284. doi:10.2337/dc10-0556.
- ^ Turnbaugh, Peter J.; Bäckhed, Fredrik; Fulton, Lucinda; Gordon, Jeffrey I. (April 2008). "Diet-Induced Obesity Is Linked to Marked but Reversible Alterations in the Mouse Distal Gut Microbiome". Cell Host & Microbe. 3 (4): 213–223. doi:10.1016/j.chom.2008.02.015.
- ^ Jump up to:a b Drake, MJ; Morris, N; Apostolidis, A; Rahnama'i, MS; Marchesi, JR (April 2017). "The urinary microbiome and its contribution to lower urinary tract symptoms". Neurourology and Urodynamics. 36 (4): 850–853. doi:10.1002/nau.23006. PMID 28444712.
- ^ Jump up to:a b Aragón, IM; Herrera-Imbroda, B; Queipo-Ortuño, MI; Castillo, E; Del Moral, JS; Gómez-Millán, J; Yucel, G; Lara, MF (January 2018). "The Urinary Tract Microbiome in Health and Disease". European Urology Focus. 4 (1): 128–138. doi:10.1016/j.euf.2016.11.001. PMID 28753805.
- ^ Jump up to:a b Schmiemann, G; Kniehl, E; Gebhardt, K; Matejczyk, M. M; Hummers-Pradier, E (2010). "The Diagnosis of Urinary Tract Infection: A Systematic Review". Deutsches Aerzteblatt Online. 107 (21): 361–367. doi:10.3238/arztebl.2010.0361. PMC 2883276.
- ^ Jump up to:a b Petrova, Mariya I.; Lievens, Elke; Malik, Shweta; Imholz, Nicole; Lebeer, Sarah (2015). "Lactobacillus species as biomarkers and agents that can promote various aspects of vaginal health". Frontiers in Physiology. 6. doi:10.3389/fphys.2015.00081.
- ^ Jump up to:a b c d Witkin, S. S.; Linhares, I. M.; Giraldo, P. (2007). "Bacterial flora of the female genital tract: Function and immune regulation". Best Practice & Research Clinical Obstetrics & Gynaecology. 21 (3): 347–354. doi:10.1016/j.bpobgyn.2006.12.004. PMID 17215167.
- ^ Todar, Kenneth (2012). "The Normal Bacterial Flora of Humans". Todar's Online Textbook of Bacteriology. Madison, WI: Kenneth Todar. Retrieved 6 April 2012.
- ^ Onderdonk, A. B.; Zamarchi, G. R.; Walsh, J. A.; Mellor, R. D.; Muñoz, A.; Kass, E. H. (1986). "Methods for quantitative and qualitative evaluation of vaginal microflora during menstruation". Applied and Environmental Microbiology. 51 (2): 333–339. PMC 238869. PMID 3954346.
- ^ Antonio, M. A. D.; Hawes, S. E.; Hillier, S. L. (1999). "The Identification of VaginalLactobacillusSpecies and the Demographic and Microbiologic Characteristics of Women Colonized by These Species". The Journal of Infectious Diseases. 180 (6): 1950–1956. doi:10.1086/315109. PMID 10558952.
- ^ Fox, Chelsea; Eichelberger, Kacey (2015). "Maternal microbiome and pregnancy outcomes". Fertility and Sterility. 104 (6): 1358–1363. doi:10.1016/j.fertnstert.2015.09.037. PMID 26493119.
- ^ Wassenaar, T.M.; Panigrahi, P. (2014). "Is a foetus developing in a sterile environment?". Letters in Applied Microbiology. 59 (6): 572–579. doi:10.1111/lam.12334. PMID 25273890.
- ^ Schwiertz, Andreas (2016). Microbiota of the human body : implications in health and disease. Switzerland: Springer. p. 1. ISBN 978-3-319-31248-4.
- ^ Franasiak, Jason M.; Scott, Richard T. (2015). "Reproductive tract microbiome in assisted reproductive technologies". Fertility and Sterility. 104 (6): 1364–1371. doi:10.1016/j.fertnstert.2015.10.012. PMID 26597628.
- ^ Sutter, V. L. (1984). "Anaerobes as normal oral flora". Reviews of infectious diseases. 6 Suppl 1: S62–S66. doi:10.1093/clinids/6.Supplement_1.S62. PMID 6372039.
- ^ Jump up to:a b c d e Purnima, Kumar (December 2013). "Oral microbiota and systemic disease". Anaerobe. 24: 90–93. doi:10.1016/j.anaerobe.2013.09.010. Retrieved 19 November 2017.
- ^ Arweiler, Nicole B.; Netuschil, Lutz (May 2016). "The Oral Microbiota". In Andreas Schwiertz. Microbiota of the Human Body: Implications in Health and Disease. Springer,Cham. pp. 45–60. ISBN 978-3-319-31248-4.
- ^ Jump up to:a b c d Avila, Maria; Ojcius, David M.; Yilmaz, Özlem (2009). "The Oral Microbiota: Living with a Permanent Guest". DNA and Cell Biology. 28 (8): 405–411. doi:10.1089/dna.2009.0874. ISSN 1044-5498.
- ^ Jump up to:a b Rogers A H (editor). (2008). Molecular Oral Microbiology. Caister Academic Press. ISBN 978-1-904455-24-0.
- ^ Jump up to:a b c d e Zarco, MF; Vess, TJ; Ginsburg, GS (2012). "The oral microbiome in health and disease and the potential impact on personalized dental medicine". Oral Diseases. 18 (2): 109–120. doi:10.1111/j.1601-0825.2011.01851.x. ISSN 1354-523X.
- ^ Man WH, de Steenhuijsen Piters WAA, Bogaert D (2017). "The microbiota of the respiratory tract: gatekeeper to respiratory health". Nature Reviews Microbiology. 15 (5): 259–270. doi:10.1038/nrmicro.2017.14. PMID 28316330.
- ^ Jump up to:a b Beringer, P M; M D Appleman (November 2000). "Unusual respiratory bacterial flora in cystic fibrosis: microbiologic and clinical features" (PDF). Current Opinion in Pulmonary Medicine. 6 (6): 545–550. doi:10.1097/00063198-200011000-00015. PMID 11100967. Archived from the original (PDF) on 16 October 2013.
- ^ Verdier, J; Luedde, T; Sellge, G (June 2015). "Biliary Mucosal Barrier and Microbiome". Viszeralmedizin. 31 (3): 156–61. doi:10.1159/000431071. PMC 4569210. PMID 26468308.
- ^ Honda, Kenya; Littman, Dan R. (7 July 2016). "The microbiota in adaptive immune homeostasis and disease". Nature. 535 (7610): 75–84. Bibcode:2016Natur.535...75H. doi:10.1038/nature18848. PMID 27383982.
- ^ Jump up to:a b c d e f g h Garrett, WS (3 April 2015). "Cancer and the microbiota". Science. 348(6230): 80–6. Bibcode:2015Sci...348...80G. doi:10.1126/science.aaa4972. PMC 5535753. PMID 25838377.
- ^ Jump up to:a b c d Gagnière, J; Raisch, J; Veziant, J; Barnich, N; Bonnet, R; Buc, E; Bringer, M. A.; Pezet, D; Bonnet, M (2016). "Gut microbiota imbalance and colorectal cancer". World Journal of Gastroenterology. 22 (2): 501–518. doi:10.3748/wjg.v22.i2.501. PMC 4716055. PMID 26811603.
- ^ Jump up to:a b Sartor, R. Balfour; Mazmanian, Sarkis K. (July 2012). "Intestinal Microbes in Inflammatory Bowel Diseases". The American Journal of Gastroenterology Supplements. 1(1): 15–21. doi:10.1038/ajgsup.2012.4.
- ^ Hold, Georgina L; Smith, Megan; Grange, Charlie; Watt, Euan Robert; El-Omar, Emad M; Mukhopadhya, Indrani (7 February 2014). "Role of the gut microbiota in inflammatory bowel disease pathogenesis: What have we learnt in the past 10 years?". World Journal of Gastroenterology. 20 (5): 1192–1210. doi:10.3748/wjg.v20.i5.1192. PMC 3921503. PMID 24574795.
- ^ Jump up to:a b Zilberman-Schapira, Gili; Zmora, Niv; Itav, Shlomik; Bashiardes, Stavros; Elinav, Hila; Elinav, Eran (2016). "The gut microbiome in human immunodeficiency virus infection". BMC Medicine. 14 (1): 83. doi:10.1186/s12916-016-0625-3. PMC 4891875. PMID 27256449.
- ^ Petrova, Mariya I.; van den Broek, Marianne; Balzarini, Jan; Vanderleyden, Jos; Lebeer, Sarah (1 September 2013). "Vaginal microbiota and its role in HIV transmission and infection". FEMS Microbiology Reviews. 37 (5): 762–792. doi:10.1111/1574-6976.12029.
- ^ Christopher Intagliata (22 December 2016). ""Necrobiome" Reveals a Corpse's Time of Death". Scientific American. Retrieved 26 March 2018.
- ^ Ed Young (10 December 2015). "Meet the Necrobiome: The Waves of Microbes That Will Eat Your Corpse". The Atlantic. Retrieved 26 March 2018.
- ^ Katherine Harmon (16 December 2009). "Bugs Inside: What Happens When the Microbes That Keep Us Healthy Disappear?". Scientific American. Retrieved 27 December 2008.
- ^ Jump up to:a b Vanagay, Pajau; et al. (1 November 2018). "US immigration westernizes the human gut microbiome" (PDF). Cell. 175: 962–972.
The Secret World Inside You Exhibit 2015-2016, American Museum of Natural History
FAQ: Human Microbiome, January 2014 American Society For Microbiology
show
v
t
e
Human microbiota
show
v
t
e
Microbiology: Bacteria
 Biology portal
Biology portal Medicine portal
Medicine portal